Welcome to the Vollmer Lab @ Northeastern University
We study the evolution and ecology of marine organisms using cutting-edge genomics techniques to ask how marine organisms respond and adapt to changing environments. Our research focuses on understanding how tropical reef-building corals are responding to emerging disease outbreaks and the global threat of a warming ocean. We combined field and tank-based experimental research with genomics to identify coral pathogens, identify genes linked to coral disease resistance, and profile the interaction between corals and their pathogens.
Research Projects
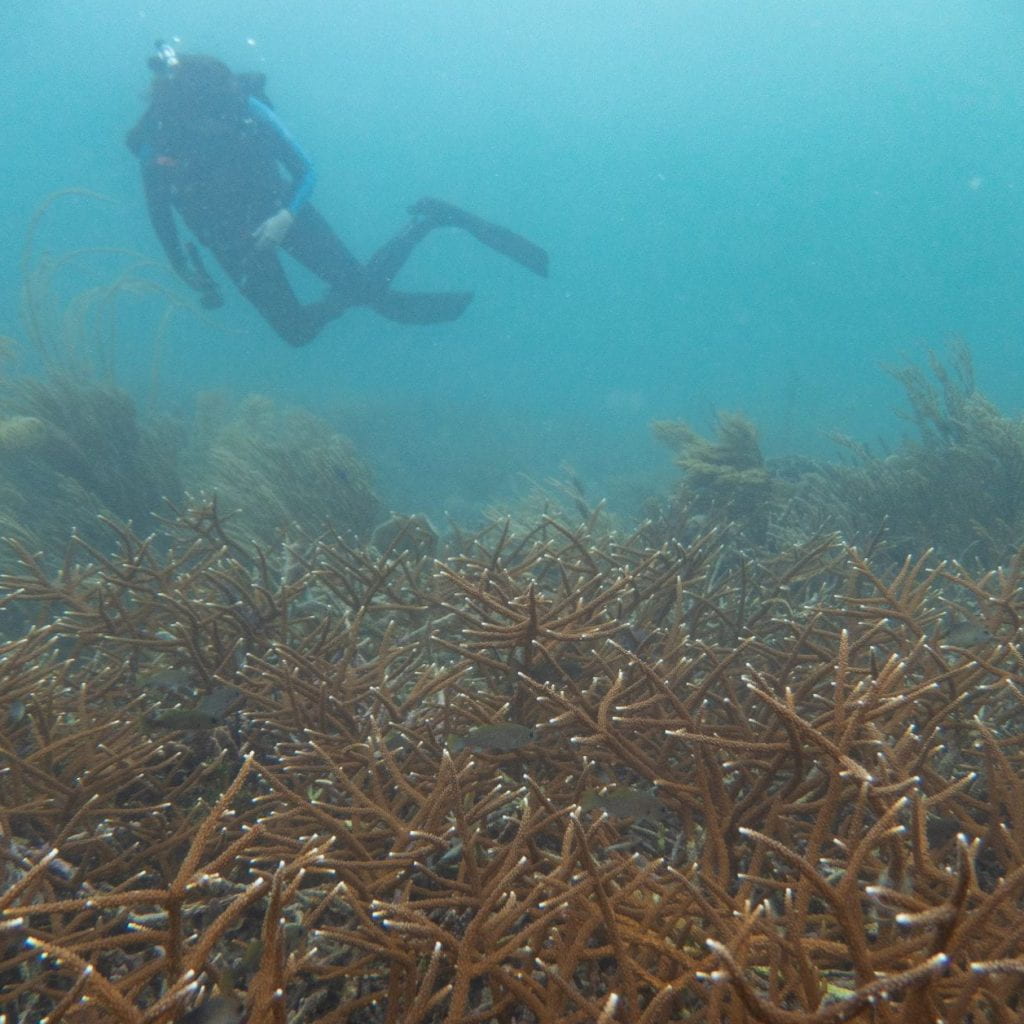

Genomics of Coral Immunity and Disease Resistance
Coral diseases, like White Band Disease and now Stony Coral Tissue Loss Disease, have decimated Caribbean reefs. Using genome-wide association analyses, our research shows that disease resistance in the Caribbean staghorn coral Acropora cervicornis has a polygenic basis and as few as 10 genes can identify resistant and susceptible corals in the wild and in coral nurseries.
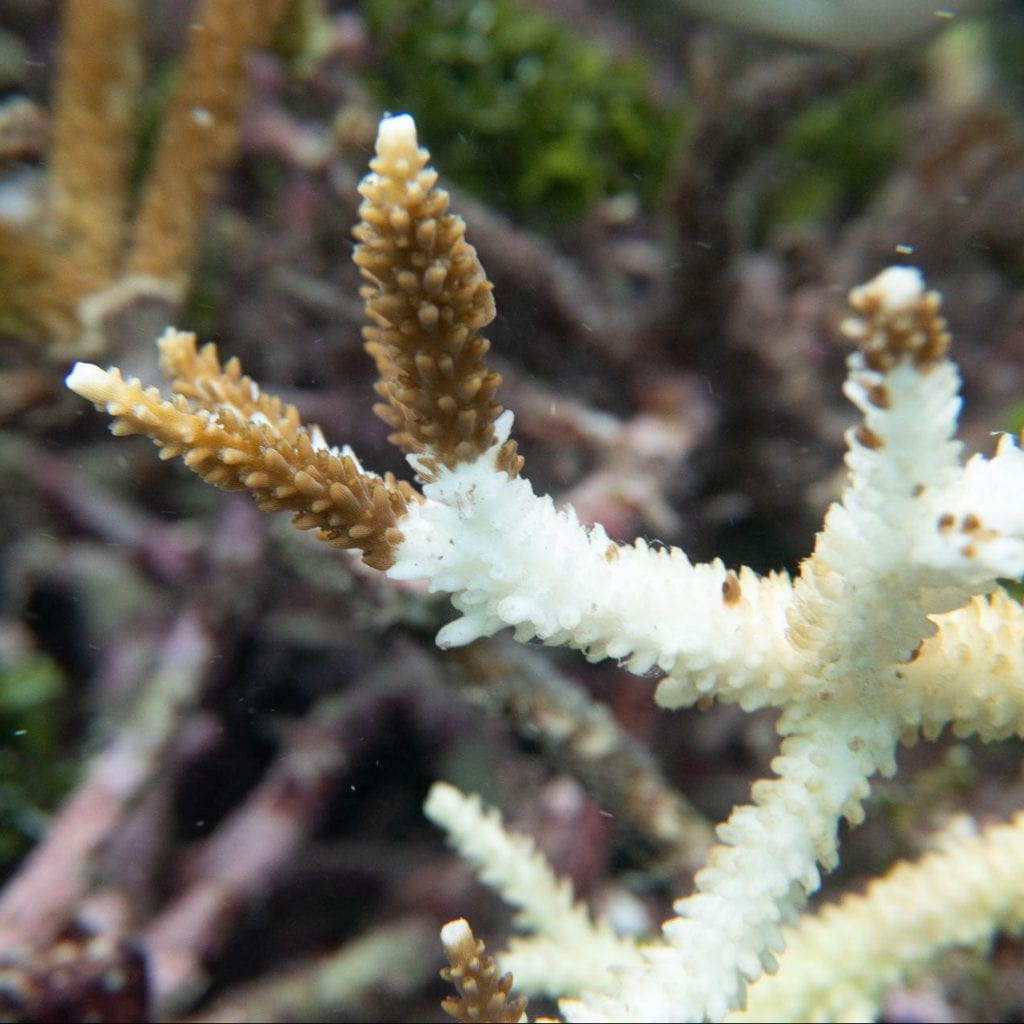

Coral Disease Etiology and Ecology
We often lack basic information about the etiology and ecology of most coral diseases. We manipulate coral microbiomes in field and tank-based experiments to hone in on primary coral pathogens and research coral-microbial interactions. We have used antibiotics and quorum sensing molecules to manipulate disease transmission and coral microbiomes to better understand who is a pathogen vs a beneficial coral bacterium.
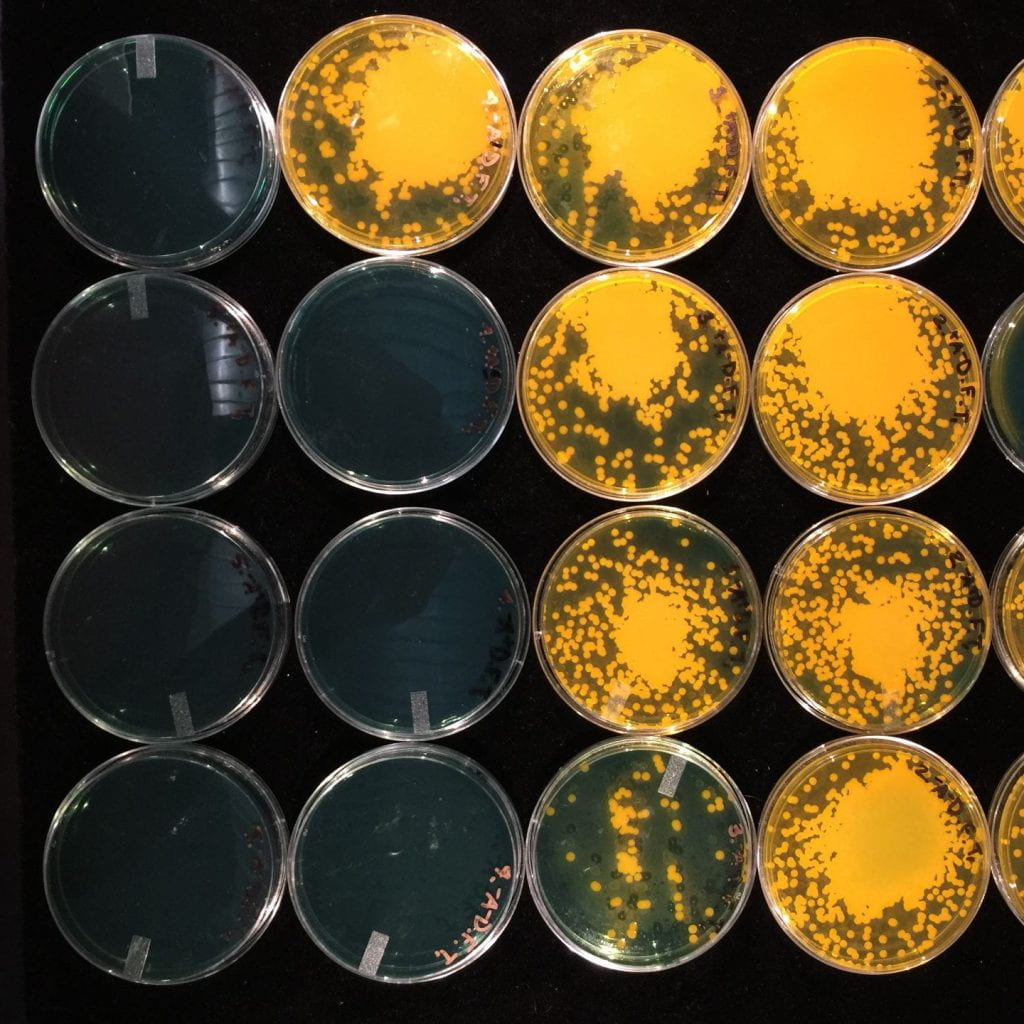
Coral-Microbial Interactions
Corals possess diverse microbiomes filled with potentially beneficial and pathogenic bacteria. Different corals have stark differences in their microbial communities and these species specific differences are stable across time and space. We aim to understand how diverse coral microbiomes structure themselves across evolutionary lineages and in space and time.
Contact: Steve Vollmer for more information about the lab